Research Interests
Evolution of the Genetic Code
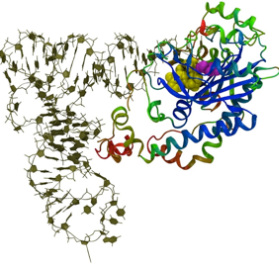
The genetic code, which is used to decode all the information in the DNA, is fundamental to all life on earth and is governed by a set of enzymes called the aminoacyl-tRNA synthetases. The evolution of the genetic code is often considered to be a frozen accident even though a few studies have shown that the genetic code may have evolved to reduce drastic effects of translational errors. The recognition of the correct pair of amino acid and tRNA molecules among a vast number of erroneous combinations is essential for the integrity of the complex cells that we observe today. I am interested in investigating the evolution of the genetic code into its present form and the implications of the formation of a robust genetic code in cells.
Robustness and Evolvability of Gene Regulatory Networks
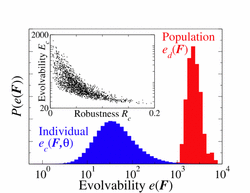
The gene regulatory networks inside a cell specify the genes that need to be expressed in a time and space dependent manner. The inference of gene regulatory networks from genomic datasets and the mechanisms by which they function remain an open question. In addition, the variation in the organization and topology of gene regulatory networks is necessary to understand the recent evolutionary history of species. I am interested in developing methods to infer the parameters of gene regulatory networks from functional genomic datasets and also studying the interplay between evolvability and robustness of these networks. In addition, I would also like to understand the variation in gene regulatory networks and prediction of phenotype based upon variation in the regulatory network inside a cell.
Variation in Enhancer Usage
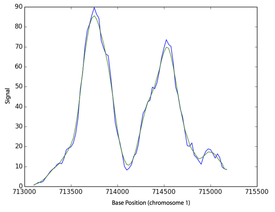
The lower cost of next generation sequencing (NGS) and the development of various functional assays involving NGS implies that genomics will be the driving force for the increase in biological knowledge during the 21st century. I am particularly interested in how these datasets can be used to understand the regulation and variation in gene expression within and between different biological species. To this end, I am developing methods to predict cis regulatory modules (elements in the genome that affect transcriptional level of genes) and methods to infer gene regulatory networks from various epigenetic and genetic datasets.
Cell type specific signaling cascades
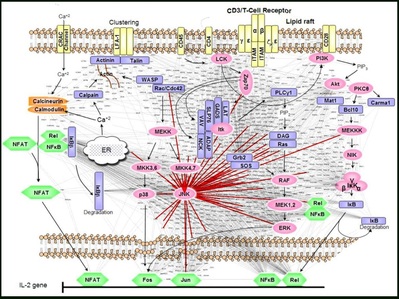
The cells ability to perceive and respond to its environment is crucial to its survival. Extracellular conditions are communicated to the cell via signaling pathways, which also coordinate the cell's response to the environmental conditions. Generally,a signaling event begins close to the cellular membrane and this signal gets amplified and transferred over large distances and timescales in a series of steps, eventually leading to changes in the transcription levels of different genes (the amount of output from information pathways) in the nucleus of the cell. The protein protein interactions is highly regulated in time (developmental stage) and space (tissue) and an understanding of how these interactions are regulated will help explain how such diverse cell types are formed from cells with identical genetic information.
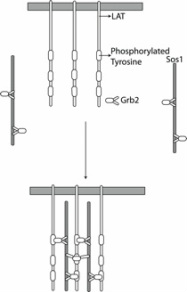
Multiscale Modeling of Genetic Regulatory Networks
In order to process the emergent behavior of biological pathways, we need to develop methods to investigate the dynamics of these processes at different resolutions in an hierarchical fashion. I want to develop methods that interface between all atom or coarse grained simulations and systems biology models of gene expression pathways inside the cell. These simulations will help predicting the phenotype given the genotype of the cell. I have already interfaced between information from all-atom molecular dynamics simulations and thermodynamic modeling to predict the stoichiometry of multivalent signaling complexes.
In order to process the emergent behavior of biological pathways, we need to develop methods to investigate the dynamics of these processes at different resolutions in an hierarchical fashion. I want to develop methods that interface between all atom or coarse grained simulations and systems biology models of gene expression pathways inside the cell. These simulations will help predicting the phenotype given the genotype of the cell. I have already interfaced between information from all-atom molecular dynamics simulations and thermodynamic modeling to predict the stoichiometry of multivalent signaling complexes.
